|
Today, investigators analyze DNA at a standardized set of more than 20 STRs. RFLP analysis has mostly been replaced by modern methods that are more precise and more sensitive–they can be done with very little DNA input. For identifying a suspect from their DNA at a crime scene, you would expect a perfect match for all the bands for a test like paternity, you would expect that at least half of the bands match between a parent and a child. Because VNTRs are so variable, the chance that two people’s banding patterns on a gel like this will match identically is extremely low. In this approach, the investigator will use several different probes, simultaneously testing several different VNTRs, and so each lane that ran on the gel will contain many visible bands, typically two (one from each chromosome) for every VNTR tested. The probes are labeled, usually with a radioactive element, allowing the investigator to create an image of where on the membrane the probes bind. These probes are DNA sequences that match specific VNTRs and will bind to the repeats that have the same sequence. The investigator then washes the membrane with DNA probes. Using a process known as southern blotting, the DNA that had been separated by size is then transferred and bound to a nylon membrane. They then take these DNA fragments and run them on a polyacrylamide gel, which separates all of the DNA fragments by size. In this procedure, an investigator takes a DNA sample isolated directly from blood or other tissue and cuts all the DNA in the sample into small pieces using an enzyme known as a restriction enzyme. The classic image of a DNA fingerprint comes from looking at VNTRs in a process known as restriction fragment length polymorphism (RFLP) analysis. 
In an STR or VNTR, the repeated sequence may occur just a few or even hundreds of times, depending on the specific locus and the individual. Repeats made of longer core sequences are referred to as variable nucleotide tandem repeats or VNTRs (also called minisatellites). When the length of the individual repeated sequence is short, usually less than 6 base pairs, these segments are called short tandem repeats or STRs (also called microsatellites). Many places in your DNA, there are regions where the same core sequence of DNA nucleotides is repeated several times over and over again, but the exact number of times the sequence is repeated can vary between individuals. The variable regions most often used in DNA fingerprinting are specific DNA segments that contain repeated strings of nucleobases. For example, can you tell which suspect (1,2,3 or 4) matches sample from the crime scene? To find a match, whether it be for a paternity test or to solve a crime, an investigator simply looks for matching patterns. The most classic protocol to visualize these differences gives the distinctive banding pattern that you have probably seen in crime TV shows and that is illustrated here. Many of these differences are in regions of DNA that contain repeated sequences known as microsatellites. To identify differences in people, researchers look at regions of DNA that are known to vary from person to person. Researchers first came up with a way to do this in the mid-1980s, and more efficient and precise methods have been developed since. To use DNA as an identification tool, researchers must first obtain quality DNA samples, know where in the DNA to look for identifying differences, and create an efficient, consistent way of looking. But of course, obtaining a DNA fingerprint is a far different process than getting someone’s actual fingerprints. DNA fingerprinting is a method for finding unique patterns in a person’s DNA and using those patterns to identify people, similar to how your unique fingerprint patterns can identify you. 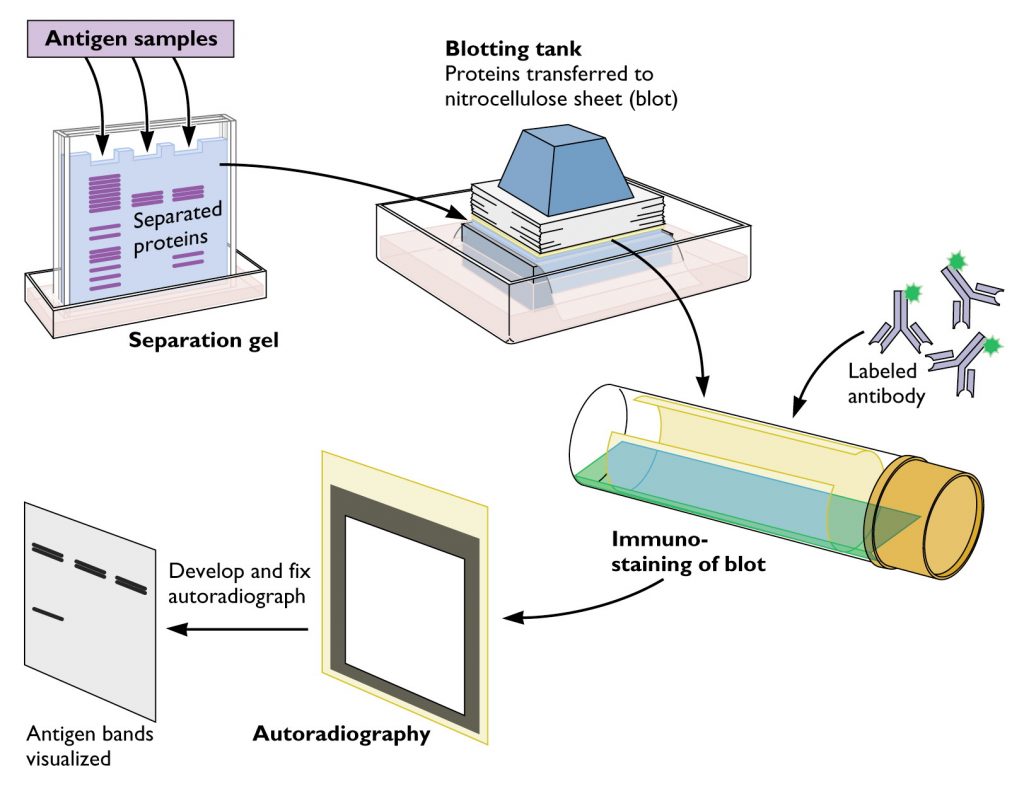
Like your fingerprints, your DNA is unique to you, doesn’t change throughout your lifetime, and whether you intend it or not, is often left behind wherever you go.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
